Clustering Pandemic COVID-19 and Relationship to Temperature and Relative Humidity Among the Tropic and Subtropic Region
DOI:
https://doi.org/10.48048/wjst.2021.9750Keywords:
COVID-19, Weather, World, Tropic, SubtropicAbstract
The outbreak of Novel Corona Virus (COVID-19) has been spreading almost in all countries of the world and become a deadly pandemic. The infections and deaths vary from high in some countries and low in others. The weather conditions significantly affect life, including viruses. In low temperature and humidity the spreading of coronavirus is expected to be fast and massive, and on the other hand, high temperature and humidity decreases the virus. However, recent data of COVID-19 shows that in tropical region infection and deaths vary of which there is a need of thorough spreading analysis. The clustering of infections and mortality at the beginning of COVID-19 outbreak was group based on the country’s profile similarity, and associated with the meteorological factors. The result shows that countries such as China, Spain, Italy and the United States have very severe attacks of COVID-19 infection. Furthermore, countries with the potential real threats of COVID-19 infections are Austria, Australia, Azerbaijan, Belgium, Bahrain, Brazil, Belarus, Canada, Switzerland, Czech Germany, Denmark, Dominican Republic, Algeria, Ecuador, Estonia, Egypt, Finland, France, Georgia, Croatia, Indonesia, Ireland, Israel, India, Iraq, Iran, Japan, Cambodia, South Korea, Kuwait, Lebanon, Sri Lanka, Lithuania, Monaco, Macedonia, Mexico, Malaysia, Nigeria, Netherlands, Norway, Nepal, New Zealand , Oman, Philippines, Pakistan, Qatar, Romania, Russia, Sweden, Singapore and Thailand. The threat of COVID-19 is not only in dry and humid sub-tropical countries, but it cannot be undermined the effect to some warm and humid tropical countries such as Brazil, Ecuador, Indonesia, Malaysia and the Philippines, which are massively infected, and the mortality rate compared to the population are very high. The study also found that dynamic humidity is a factor that must be considered, especially in the tropics.
HIGHLIGHTS
- The COVID-19 pandemic that originated in Wuhan, China spreads rapidly around the world
- Demographics and weather are thought to influence the spreading and death of COVID-19
- Clustering of demographic and weather factors on COVID-19 shows that countries such as China, Spain, Italy, and the United States are experiencing severe attacks of COVID-19 infection
- Covid threatens countries with high population density or large populations
- Although warm and humid temperatures in the tropics such as Brazil, Ecuador, Indonesia, Malaysia, and the Philippines can a little slow the spreading of infection, the risk of COVID-19 infection remains high
GRAPHICAL ABSTRACT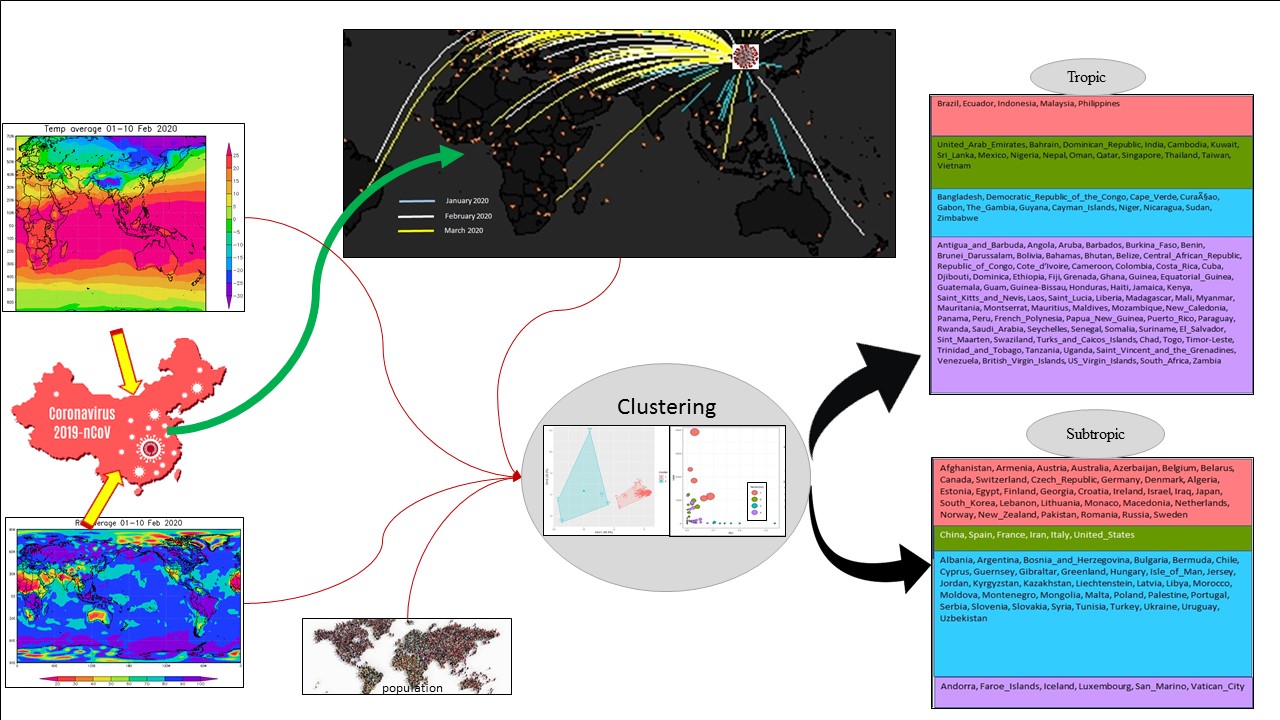
Downloads
Metrics
References
N Zhu, Z Zhang, W Wang, X Li, B Yang, J Song, X Zhao, B Huang, W Shi, R Lu, P Niu and F Zhan. A novel coronavirus from patients with pneumonia in China, 2019. N. Engl. J. Med. 2020; 382, 727-33.
C Huang, Y Wang, X Li, L Ren, J Zhao, Y Hu, L Zhang, G Fan, J Xu, X Gu, Z Cheng, T Yu, J Xia, Y Wei, W Wu, X Xie, W Yin, H Li, M Liu, Y Xiao, H Gao, L Guo, J Xie, G Wang, R Jiang, Z Gao, Q Jin, J Wang and B Cao. Clinical features of patients infected with 2019 novel coronavirus in Wuhan, China. Lancet 2020; 395, 497-506.
Coronavirus disease (COVID-19) weekly epidemiological update and weekly operational update, Available at: https://www.who.int/emergencies/diseases/novel-coronavirus-2019/situation-reports, accessed January 2020.
EL Gralinski and VD Menachery. Return of the coronavirus: 2019-ncov. Viruses 2020; 12, 135.
P Zhou, XL Yang, XG Wang, B Hu, L Zhang, W Zhang, HR Si, Y Zhu, B Li, CL Huang, HD Chen, J Chen, Y Luo, H Guo, RD Jiang, MQ Liu, Y Chen, XR Shen, X Wang, XS Zheng, K Zhao, QJ Chen, F Deng, LL Liu, B Yan, FX Zhan, YY Wang, GF Xiao and ZL Shi. A pneumonia outbreak associated with a new coronavirus of probable bat origin. Nature 2020; 579, 270-3.
D Xu, Z Zhang, F Chu, Y Li, L Jin, L Zhang, GF Gao and FS Wang. Genetic Variation of SARS Coronavirus in Beijing Hospital. Emerg. Infect. Dis. 2004; 10, 789-94.
Y Fan, K Zhao, ZL Shi and P Zhou. Bat coronaviruses in China. Viruses 2019; 11, 1-14.
EC Teeling, MS Springer, O Madsen, P Bates, SJ O’Brien and WJ Murphy, A molecular phylogeny for bats illuminates biogeography and the fossil record. Science 2005; 307, 580-4.
Worldometers: Covid-19 coronavirus pandemic, Available at: https://www.worldometers.info/ coronavirus, accessed January 2020.
KH Chan, JS MalikPeiris, SY Lam, LLM Poon, KY Yuen and WH Seto. The effects of temperature and relative humidity on the viability of the SARS coronavirus. Adv Virol. 2011; 734690, 1-7.
MER Darnell, K Subbarao, SM Feinstone and DR Taylor. Inactivation of the coronavirus that induces severe acute respiratory syndrome, SARS-CoV. J. Virol. Meth. 2004; 121, 85-91.
MK Ijaz, AH Brunner and SA Sattar. Survival characteristics of airborne human coronavirus 229E. J. Gen. Virol. 1985; 66, 2743-8.
J Sizun, MWN Yu and PJ Talbot. Survival of human coronaviruses 229E and OC43 in suspension and after drying on surfaces: a possible source of hospital-acquired infections. J. Hosp. Infect. 2000; 46, 55-60.
HF Rabenau, J Cinatl, B Morgenstern, G Bauer, W Preiser and HW Doerr. Stability and inactivation of SARS coronavirus. Med. Microbiol. Immunol. 2005; 194, 1-6.
J Tan, L Mu, J Huang, S Yu, B Chen and J Yin. An initial investigation of the association between the SARS outbreak and weather: With the view of the environmental temperature and its variation. J. Epidemiol. Commun. Health. 2005; 59, 186-92.
J Yuan, H Yun, W Lan, W Wang, SG Sullivan, S Jia and AH Bittles. A climatologic investigation of the SARS-CoV outbreak in Beijing, China. Am. J. Infect. Control. 2006; 34, 234-6.
QC Cai, J Lu, QF Xu, Q Guo, DZ Xu, QW Sun, H Yang, GM Zhao and QW Jiang. Influence of meteorological factors and air pollution on the outbreak of severe acute respiratory syndrome. Public Health 2007; 121, 258-65.
LM Casanova, S Jeon, WA Rutala, DJ Weber and MD Sobsey. Effects of air temperature and relative humidity on coronavirus survival on surfaces. Appl. Environ. Microbiol. 2010; 76, 2712-7.
GBDI Collaborators. Mortality, morbidity, and hospitalisations due to influenza lower respiratory tract infections, 2017: An analysis for the global burden of disease study 2017. Lancet. Respir. Med. 2019; 7, 69-89.
C Viboud, WJ Alonso and L Simonsen. Influenza in tropical regions. PLoS Med. 2006; 3, e89.
KB Feshbach, WJ Alonso, V Charu, J Tamerius, L Simonsen, MA Miller and C Viboud. Latitudinal variations in seasonal activity of influenza and respiratory syncytial virus (RSV): A global comparative review. PLoS One 2013; 8, e54445.
Y Li, RM Reeves, X Wang, Q Bassat, WA Brooks, C Cohen, DP Moore, M Nunes, B Rath,H Campbell and H Nair. Global patterns in monthly activity of influenza virus, respiratory syncytial virus, parainfluenza virus, and metapneumovirus: A systematic analysis. Lancet. Glob. Health 2019; 7, e1031-e45.
MM Sajadi, P Habibzadeh, A Vintzileos, S Shokouhi, FM Wilhelm and A Amoroso. Temperature and latitude analysis to predict potential spread and seasonality for covid-19, Available at: http://dx.doi.org/10.2139/ssrn.3550308, accessed January 2020.
European Centre for Disease Prevention and Control, Available at: https://www.ecdc.europa.eu, accessed January 2020.
Simplemaps, Available at: https://simplemaps.com, accessed January 2020.
Physical Sciences Laboratory, Available at: https://www.esrl.noaa.gov/psd/data/gridded/ data.ncep.reanalysis.pressure.html, accessed January 2020.
C Marzban and S Sandgathe. Cluster analysis for verification of precipitation fields. Weather Forecast. 2006; 21, 824-38.
S Gokila, KA Kumar and A Bharath. Clustering and classification in support of climatology to mine weather data: A review. Int. J. Comput. Algorithm. 2015; 4, 45-8.
B Murugesakumar, K Anandakumar and A Bharathi. Improved fuzzy K-means cluster algorithm to analyse weather data in coimbatoreregion. Int. J. Adv. Eng. Res. Dev. 2017; 4, 840-6.
W Dong, K Yang, Q Xu, L Liu and J Chen. Spatio-temporal pattern analysis forevaluation of the spread of humaninfections with avian influenza A(H7N9) virus in China, 2013-2014. MC Infect. Dis. 2017; 17, 1-17.
CR Vicente, KH Herbinger, CC Junior, CM Romano, ASA Cabidelle and G Froschl. Determination of clusters and factors associated with dengue dispersion during the first epidemic related to Dengue virus serotype4 in Vitória, Brazil. PLoS One 2016; 12, e0175432.
RM McCloskey and AY Poon. A model-based clustering method to detect infectious disease transmission out breaks from sequence variation. PLoS Comput. Biol. 2017; 13, e1005868.
C Yuan and H Yang. Research on K-value selection method of K-MeansClustering algorithm. J. Multidiscip. Sci. 2019; 2, 227-35.
JP Ortega, NNA Ortega, and D Romero. Balancing effort and benefit of K-means clustering algorithms in big data realms. PLoS One 2018; 13, 1-19.
MZ Sáska and S Dombay. Seasons's shifts in some depressions of the Eastern Carpathians, based on daily temperature analysis. In: Proceedings of the Air and Water–Components of the Environment, Cluj-Napoca, Romania. 2020, p. 213-22.
Downloads
Published
How to Cite
Issue
Section
License
Copyright (c) 2021 Walailak University

This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License.