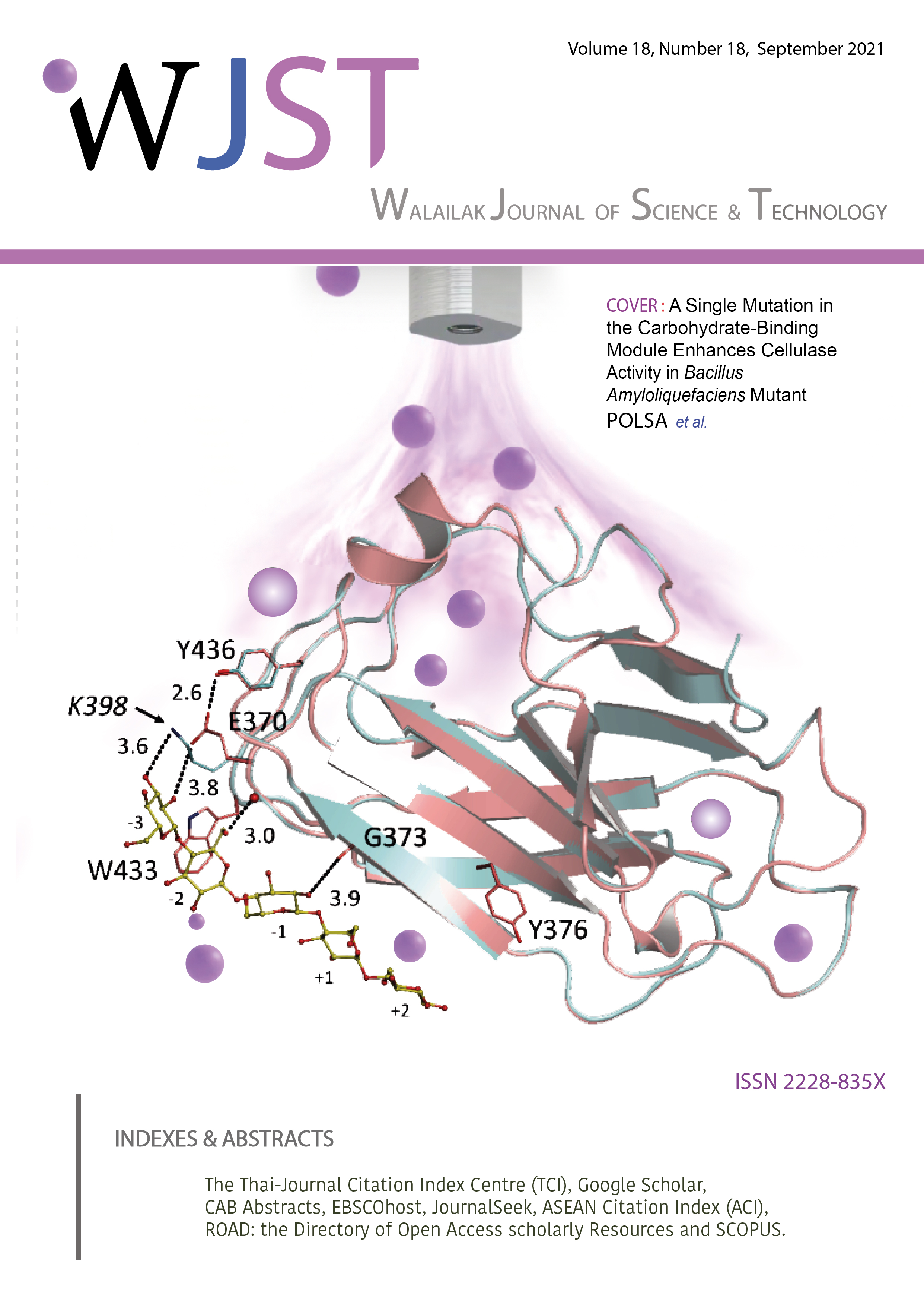
A Single Mutation in the Carbohydrate-Binding Module Enhances Cellulase Activity in Bacillus Amyloliquefaciens Mutant
DOI:
https://doi.org/10.48048/wjst.2021.23985Keywords:
Agricultural wastes, Bacillus amyloliquefaciens, Carbohydrate-binding module (CBM), Cellulase, MutationAbstract
From our earlier work, we modified the carbohydrate-binding module (CBM) of Bacillus amyloliquefaciens to increase cellulase activity using cold plasma technology. The cellulase gene (BglC-M) from the mutant was expressed in Escherichia coli BL21(DE3) under the T7 promoter. The hydrolysis activity of the cellulase mutant (BglC-M) was approximately 2.5-fold higher than the control (BglC-W) over a wide range of pH and temperature conditions. The amino acid sequence of the mutant BglC-M contained 471 residues that were almost identical to the control BglC-W. Only a single amino acid, lysine, was replaced by glutamic acid at position 370 (K370E) within the carbohydrate-binding module (CBM). Structure prediction and substrate docking of BglC-M indicated that the single mutation (K370E) might involve cellulose binding of the β-sandwich facilitated by hydrogen bonding. The docking study of cellopentaose with the model structure of BglC-M indicated that the replacement of lysine-370 led to the formation of a hydrogen bond with 436Y, which has a shorter distance (2.6 Å) compared with the control (5.4 Å). As a result, the structure becomes more compact and stable, resulting in increased catalytic efficiency. Finally, the biomass hydrolysis ability of cellulase was investigated on lignocellulosic wastes such as pineapple peel, corncob, and durian peel. The BglC-M enzyme showed a more significant amount of reducing sugar released from all lignocellulosic wastes than the control. This was the first evidence that altering the base composition of the cellulose binding module enhanced the catalytic activity.
HIGHLIGHTS
- Increasing cellulase activity of Bacillus amyloliquefaciens using plasma technology
- Mutation at cellulose-binding module enhance cellulase hydrolysis activity
- Greater cellulase activity in the hydrolysis on lignocellulosic wastes
GRAPHICAL ABSTRACT
Downloads
Metrics
References
T Hasunuma, F Okazaki, N Okai, KY Hara, J Ishii and A Kondo, A review of enzymes and microbes for lignocellulosic biorefinery and the possibility of their application to consolidated bioprocessing technology. Bioresour. Technol. 2013; 153, 513-22.
AM Kholif, MA El-Ashry, HA El-Alamy, HM El-Sayed, M Fadel and SM Kholif. Biological treatments banana wastes for feeding lactating goats. Egypt. J. Nutr. Feeds 2005; 8, 149-62.
J Pérez, J Muñoz-Dorado, T de la Rubia and J Martínez, Biodegradation and biological treatments of cellulose, hemicellulose, and lignin: An overview. Int. Microbiol. 2002; 5, 53-63.
LT Angenent, K Karim, MH Al-Dahlan, BA Wrenn and R Domiquez-Espinosa. Production of bioenergy and biochemicals from industrial and agricultural wastewater. Trends. Biotechnol. 2004; 22, 477-85.
HH Azzaz. 2009, Effect of cellulytic enzymes addition to diets on the productive performance of lactating goats. Master Thesis. Cairo University, Cairo, Egypt.
J Hong, H Tamaki, S Akiba, K Yamamoto and H Kumagai. Cloning of a gene encoding a highly stable endo-beta-1,4-glucanase from Aspergillus niger and its expression in yeast. J. Biosci. Bioeng. 2001; 92, 434-41.
YH Li, M Ding, J Wang, GJ Xu and F Zhao. A novel thermoacidophilic endoglucanase, Ba-EGA, from a new cellulose-degrading bacterium, Bacillus sp. AC-1. Appl. Microbiol. Biotechnol. 2006; 70, 430-6.
M Kurasin and P Väljamäe. Processivity of cellobiohydrolases is limited by the substrate. J. Biol. Chem. 2011; 286, 169-77.
AB Boraston, DN Bolam, HJ Gilbert and GJ Davies. Carbohydrate-binding modules: Fine-tuning polysaccharide recognition. Biochem. J. 2004; 382, 769-81.
V Lombard, H Golaconda Ramulu, E Drula, PM Coutinho and B Henrissat. The carbohydrate-active enzymes database (CAZy) in 2013. Nucleic. Acids Res. 2014; 42, D490-D495.
R Du, S Jiao, Y Dai, J An, J Lv, X Yan, J Wang and B Han. Probiotic bacillus amyloliquefaciens C-1 improves growth performance, stimulates GH/IGF-1, and regulates the gut microbiota of growth-retarded beef calves. Front. Microbiol. 2018; 9, 2006.
OS Kotchoni, O Shonukan and WE Gachomo. Bacillus pumilus BpCRI 6 a promising candidate for cellulase production under conditions of catabolite repression. Afr. J. Biotechnol. 2003; 2, 140-6.
M Mandels, J Weber and R Parizek. Enhanced cellulase production by a mutant of Trichoderma viride. Appl. Microbiol. 1971; 21, 152-4.
DE Eveleigh and BS Montenecourt. Increasing yields of extracellular enzymes. Adv. Appl. Microbiol. 1979; 25, 57-74.
G Liu, L Zhang, Y Qin, G Zou, Z Li, X Yan, X Wei, M Chen, L Chen, K Zheng, J Zhang, L Ma, J Li, R Liu, H Xu, X Bao, X Fang, L Wang, Y Zhong, W Liu, H Zheng, S Wang, C Wang, L Xun, GP Zhao, T Wang, Z Zhou and Y Qu. Long-term strain improvements accumulate mutations in regulatory elements responsible for hyper-production of cellulolytic enzymes. Sci. Rep. 2013; 3, 1569.
SA Mahadevan, SG Wi, DS Lee and HJ Bae. Site-directed mutagenesis and CBM engineering of Cel5A (Thermotoga maritima). FEMS. Microbiol. Lett. 2008; 287, 205-11.
N Polsa, W Suyotha, S Suebsan, S Anuntalabhochai and K Sangwijit. Increasing xylanase activity of Bacillus subtilis by atmospheric pressure plasma jet for biomass hydrolysis. 3 Biotech. 2020; 10, 22.
SP Voutilainen, T Puranen, M Siika-Aho, A Lappalainen, M Alapuranen, J Kallio, S Hooman, L Viikari, J Vehmaanperä and A Koivula. Cloning, expression, and characterization of novel thermostable family 7 cellobiohydrolases. Biotechnol. Bioeng. 2008; 101, 515-28.
AV Gusakov. Alternatives to Trichoderma reesei in biofuel production. Trends Biotechnol. 2011; 29, 419-25.
T Lengauer and M Rarey. Computational methods for biomolecular docking. Curr. Opin. Struct. Biol. 1996; 6, 402-6.
K Naito, M Kusaba, N Shikazono, T Takano, A Tanaka, T Tanisaka and M Nishimura. Transmissible and nontransmissible mutations induced by irradiating Arabidopsis thaliana pollen with gamma-rays and carbon ions. Genetics 2005; 169, 881-9.
KP Gopinath, S Murugesan, J Abraham and K Muthukumar. Bacillus sp. mutant for improved biodegradation of congo red: Random mutagenesis approach. Bioresour. Technol. 2009; 100, 6295-300.
K Sangwijit, J Jitonnom, S Pitakrattananukool, LD Yu and S Anuntalabhochai. Low-energy plasma immersion ion implantation modification of bacteria to enhance hydrolysis of biomass materials. Surf. Coat. Technol. 2016; 306, 336-40.
F Islam and N Roy. Screening purification and characterization of cellulase from cellulase producing bacteria in molasses. BMC Res. Notes 2018; 11, 1-6.
GC Miller. Use of dinitrosalicylic acid reagent for determination of reducing sugar. Anal. Chem. 1959; 31, 426-8.
MA Larkin, G Blackshields, NP Brown, R Chenna, PA McGettigan, H McWilliam, F Valentin, IM Wallace, A Wilm, R Lopez, JD Thompson, TJ Gibson and DG Higgins. Clustal W and Clustal X version 2.0. Bioinformatics 2007; 23, 2947-8.
X Robert and P Gouet. Deciphering key features in protein structures with the new ENDscript server. Nucleic. Acids Res. 2014; 42, W320-W324.
A Waterhouse, M Bertoni, S Bienert, G Studer, G Tauriello, R Gumienny, FT Heer, TAP de Beer, C Rempfer, L Bordoli, R Lepore and T Schwede. SWISS-MODEL: Homology modelling of protein structures and complexes. Nucleic. Acids Res. 2018; 46, W296-W303.
O Trott and AJ Olson, AutoDock Vina: Improving the speed and accuracy of docking with a new scoring function, efficient optimization, and multithreading. J. Comput. Chem. 2010; 31, 455-61.
The PyMOL Molecular Graphics System, Available at: http://pymol.sourceforge.net/overview/index.htm, accessed June 2019.
R Kalawong, M Wakayama, S Anuntalabhochai, C Wongsawad and K Sangwijit. Comparison and characterization of purified cellulase and xylanase from Bacillus amyloliquefaciens CX1 and Bacillus subtilis B4. Chiang. Mai. J. Sci. 2018; 45, 92-105.
P McDonald, AR Henderson and R Whittenbury. The effect of temperature on ensilage. J. Sci. Food Agric. 1966; 17, 476-80.
CR Santos, JH Paiva, ML Sforça, JL Neves, RZ Navarro, J Cota, PK Akao, ZB Hoffmam, AN Meza, JH Smetana, ML Nogueira, I Polikarpov, J Xavier-Neto, FM Squina, RJ Ward, R Ruller, AC Zeri and MT Murakami. Dissecting structure-function-stability relationships of a thermostable GH5-CBM3 cellulase from Bacillus subtilis 168. Biochem. J. 2012; 441, 95-104.
O Yaniv, S Petkun, L Shimon, B Edward, R Lamed and F Frolow. A single mutation reforms the binding activity of an adhesion-deficient family 3 carbohydrate-binding module. Acta Crystallogr. D. Biol. Crystallogr. 2012; 68, 819-28.
AB Boraston, DN Bolam, HJ Gilbert and GJ Davies. Carbohydrate-binding modules: Fine-tuning polysaccharide recognition. Biochem. J. 2004; 382, 769-81.
L Ma, R Aizhan, Wang, X Wang, Y Yi, Y Shan, B Liu, Y Zhou and X Lu. Cloning and characterization of low-temperature adapted GH5-CBM3 endo-cellulase from Bacillus subtilis 1AJ3 and their application in the saccharification of switchgrass and coffee grounds. AMB Expr. 2020; 10, 42.
KK Ghadikolaei, J Gharechahi, K Haghbeen, K Noghabi, GH Salekdeh and H Zahiri. A cold-adapted endoglucanase from camel rumen with high catalytic activity at moderate and low temperatures: An anomaly of truly cold-adapted evolution in a mesophilic environment. Extremophiles 2018; 22, 315-26.
XX Yan, XM An, LL Gui and DC Liang. From structure to function: Insights into the catalytic substrate specificity and thermostability displayed by Bacillus subtilis mannanase BCman. J. Mol. Biol. 2008; 379, 535-44.
F Contreras, S Pramanik, AM Rozhkova, N Zorov, OI Korotkova, AP Sinitsyn, U Schwaneberg, and MD Davari. Engineering robust cellulases for tailored lignocellulosic degradation cocktails. Int. J. Mol. Sci. 2020; 21, 1589.
SF Badino, SJ Christensen, J Kari, MS Windahl, S Hvidt, K Borch and P Westh. Exo-exo synergy between Cel6A and Cel7A from Hypocrea jecorina: Role of carbohydrate binding module and the endo-lytic character of the enzymes. Biotechnol. Bioeng. 2017; 114, 1639-47.
P Westh, K Borch, T Sørensen, R Tokin, J Kari, S Badino, MA Cavaleiro, N Røjel, S Christensen, CS Vesterager and C Schiano-di-Cola. Thermoactivation of a cellobiohydrolase. Biotechnol. Bioeng. 2018; 115, 831-8.
Downloads
Published
How to Cite
Issue
Section
License

This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License.