Isolation and Identification of Cellulolytic Bacteria Symbiont from Various Termites on Different Nest Type in Bukit Baka Bukit Raya National Park, West Kalimantan, Indonesia
DOI:
https://doi.org/10.48048/wjst.2021.12708Keywords:
Bacillus, Bukit Baka-Bukit Raya National Park, Endoglucanase producing bacteria, Termite intestinal tractAbstract
The microbial symbiotic community in the digestive tract of termites is reportedly influenced by the taxonomy and feeding habit of the host. Both factors are strongly correlated with the nest type. This study aimed to isolate the cellulolytic bacteria from termite’s digestive tract on different nest types and characterize and identify the potential isolates. The research methods included termite sampling conducted in Bukit Baka Bukit Raya National Park (BBBRNP), Melawi, West Kalimantan, isolation of cellulolytic bacteria from termites’ gut, endoglucanase activity test, biochemical characterization, and DNA analysis based on the amplification of 16S rRNA gene. Thirty isolates from 6 different species of termites on three different nest types were successfully isolated. Sixteen potential endoglucanase bacterial isolates were tested in terms of their endoglucanase activity. The cellulolytic index measured from those isolates ranged from 1.162 - 4.894. Three isolates (MRH.13.S, MRH.13.AF, and MRH.13.O2) with the highest cellulolytic index on each nest type were identified. The analysis of 16S rRNA gene using BLAST (Basic Local Alignment Search Tool for Nucleotides) revealed that isolate MRH.13.S had the closest relationship with Bacillus tequilensis (99 % homology). Based on biochemical characterization, MRH.13.AF and MRH.13.O2 isolates were related to Bacillus spp.
HIGHLIGHTS
- Potential cellulolytic bacteria from termite intestinal tract from different nests (i.e., soil, wood, and arboreal) were isolated and compared
- Termites were obtained from a lowland dipterocarp primary forest ecosystem in Bukit Baka Bukit Raya National Park, West Kalimantan Province, Indonesia
- Termite species collected were Termes comis, Dicuspiditermes garthwaitei, Synhamitermes quadriceps, Havilanditermes proatripennis, Bulbitermes borneensis, and Bulbitermes parapusillus
- Potential cellulolytic bacteria acquired were closely related with Bacillus tequilensis and Bacillus spp
GRAPHICAL ABSTRACT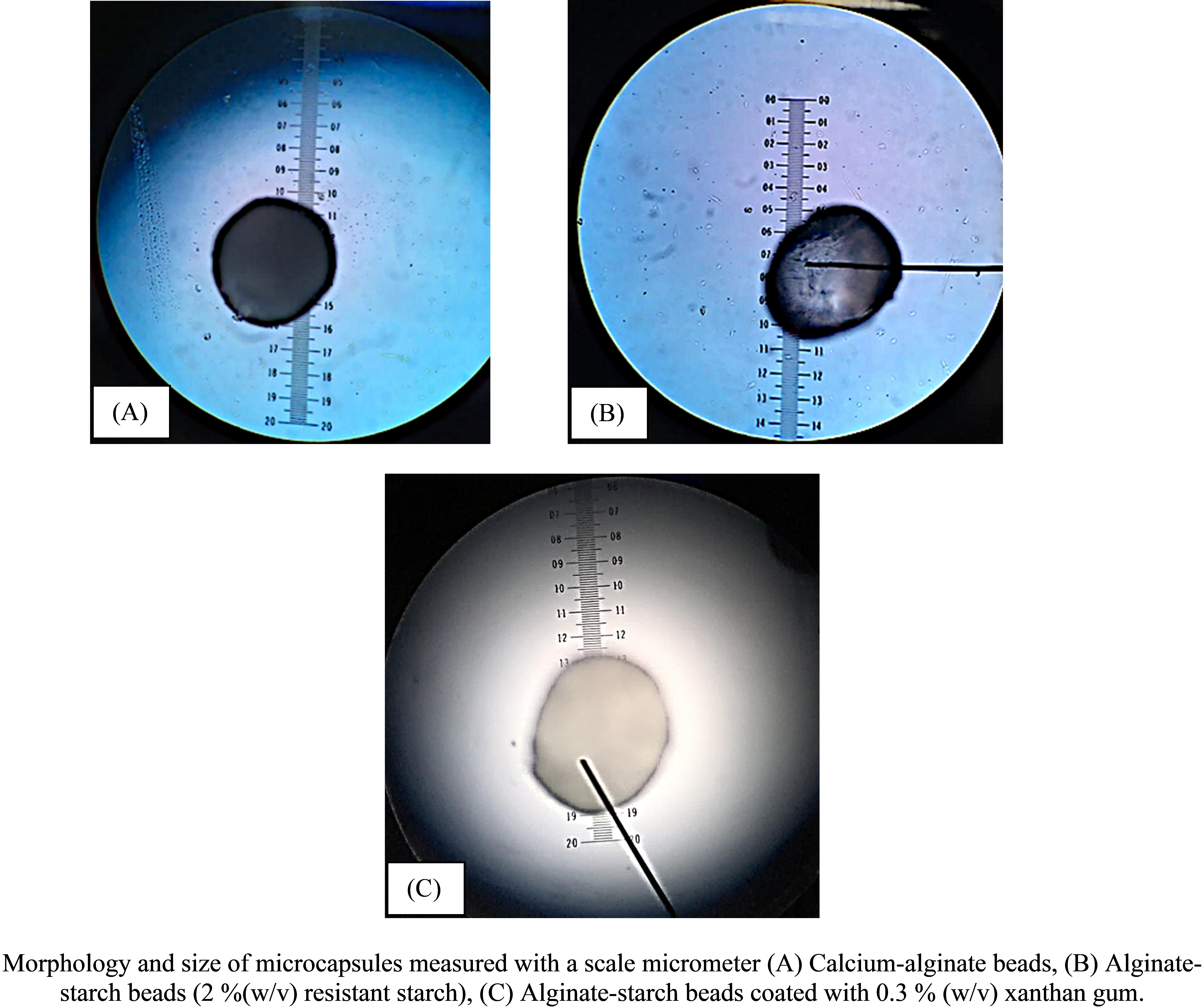
Downloads
Metrics
References
HMN Iqbal, G Kyazze and T Keshavarz. Advances in the valorization of lignocellulosic materials by biotechnology: An overview. BioResources 2013; 8, 3157-76.
Z Anwar, M Gulfraz and M Irshad. Agro-industrial lignocellulosic biomass a key to unlock the future bio-energy: A brief review. J. Radiat. Res. Appl. Sci. 2014; 7, 163-73.
M Bilal, M Asgher, HMN Iqbal, H Hu and X Zhang. Biotransformation of lignocellulosic materials into value-added products-a review. Int. J. Biol. Macromol. 2017; 98, 447-58.
G Guerriero, JF Hausman, J Strauss, H Ertan and KS Siddiqui. Lignocellulosic biomass: Biosynthesis, degradation, and industrial utilization. Eng. Life Sci. 2016; 16, 1-16.
L Auer, A Lazuka, D Sillam-Dussès, E Miambi, M O'Donohue and G Hernandez-Raquet. Uncovering the potential of termite gut microbiome for lignocellulose bioconversion in anaerobic batch bioreactors. Front. Microbiol. 2017; 8, 2623.
P Eggleton. An introduction to termites: Biology, taxonomy and functional morphology. In: DE Bignell, Y Roisin and N Lo (Eds.). Biology of termites: A modern synthesis. Springer, Dordrecht, 2010, p. 1-26.
A Thong-On, K Suzuki, S Noda, J Inoue, S Kajiwara and M Ohkuma. Isolation and characterization of anaerobic bacteria for symbiotic recycling of uric acid nitrogen in the gut of various termites. Microbes Environ. 2012; 27, 186-92.
J Reuß, R Rachel, P Kämpfer, A Rabenstein, J Küver, S Dröge and H König. Isolation of methanotrophic bacteria from termite gut. Microbiol. Res. 2015; 179, 29-37.
Z Pourramezan, G Ghezelbash, B Romani, S Ziaei and A Hedayatkhah. Screening and identification of newly isolated cellulose-degrading bacteria from the gut of xylophagous termite Microcerotermes diversus (Silvestri). Microbiologiia 2012; 81, 736-42.
D Kavitha, K Vijayarani and K Kumanan. 16S rRNA typing of cellulolytic bacteria from the termite Odontotermes formosanus. Ind. J. Vet. Anim. Sci. Res. 2014; 43, 359-68.
A Ferbiyanto, I Rusmana and R Raffiudin. Characterization and identification of cellulolytic bacteria from gut of worker Macrotermes gilvus. Hayati J. Biosci. 2015; 22, 197-200.
RB Rosengaus, JE Moustakas, DV Calleri and JFA Traniello. Nesting ecology and cuticular microbial loads in dampwood (Zootermopsis angusticollis) and drywood termites (Incisitermes minor, I. schwarzi, Cryptotermes cavifrons). J. Insect Sci. 2003; 3, 31.
A Manjula, M Pushpanathan, S Sathyavathi, P Gunasekaran and J Rajendhran. Comparative analysis of microbial diversity in termite gut and termite nest using ion sequencing. Curr. Microbiol. 2016; 72, 267-75.
Bukit Baka Bukit Raya National Park. Management portrait of Bukit Baka Bukit Raya National Park. Bukit Baka Bukit Raya National Park, Sintang, 2012, p. 3-16.
Indonesian Agency for Meteorogy Climatology and Geophysics, Daily Climate Data for West Kalimantan Province, Availabe at: http://dataonline.bmkg.go.id/ketersediaan_data, accessed December 2016.
M Ahmad. Key to the Indomalayan termites. Biologia 1958; 4, 119-8.
M Ahmad. Termites (Isoptera) of Thailand. American Museum of Natural History, New York, USA, 1965, p. 1-113.
Y Sornnuwat, C Vongkaluang and Y Takematsu. A systematic key to termites of Thailand. Kasetsart J. Nat. Sci. 2004; 38, 349-68.
M Ramin, AR Alimon and N Abdullah. Identification of cellulolytic bacteria isolated from the termite Coptotermes curvignathus (Holmgren). J. Rapid Meth. Aut. Microbiol. 2009; 17, 103-16.
RC Kasana, R Salwan, H Dhar, S Dutt and A Gulati. A rapid and easy method for the detection of microbial cellulases on agar plates using Gram’s iodine. Curr. Microbiol. 2008; 57, 503-7.
C Florencio, S Couri and CS Farinas. Correlation between agar plate screening and solid-state fermentation for the prediction of cellulase production by Trichoderma strains. Enzyme Res. 2012; 2012, 793708.
Y Nakagawa, T Sakane, M Suzuki and K Hatano. Phylogenetic structure of the genera Flexibacter, Flexithrix, and Microscilla deduced from 16S rRNA sequence analysis. J. Gen. Appl. Microbiol. 2002; 48, 155-65.
F Jeanmougin, JD Thompson, M Gouy, DG Higgins and TJ Gibson. Multiple sequence alignment with Clustal X. Trends Biochem. Sci. 1998; 23, 403-5.
SE Donovan, P Eggleton and DE Bignell. Gut content analysis and a new feeding group classification of termites. Ecol. Entomol. 2001; 26, 356-66.
CG Liu, K Li, Y Wen, BY Geng, Q Liu and YH Lin. Bioethanol: New opportunities for an ancient product. In: Y Li and X Ge (Eds.). Advances in Bioenergy. Elsevier, USA, 2019.
MA Postava-Davignon. 2010, Evolution and ecology of termite nesting behavior and its impact on disease susceptibility. Ph. D. Dissertasion. Northeastern University, Boston, USA.
T Shankar, V Mariappan and L Isaiarasu. Screening cellulolytic bacteria from the mid-gut of the popular composting earthworm Eudrilus eugeniae (Kinberg). World J. Zool. 2011; 6, 142-8.
P Sheng, S Huang, Q Wang, A Wang and H Zhang. Isolation, screening, and optimization of the fermentation conditions of highly cellulolytic bacteria from the hindgut of Holotrichia parallela larvae (Coleoptera: Scarabaeidae). Appl. Biochem. Biotechnol. 2012; 167, 270-84.
L Adams and R Boopathy. Isolation and characterization of enteric bacteria from the hindgut of Formosan termite. Bioresource Technol. 2005; 96, 1592-8.
P Gupta, K Samant and A Sahu. Isolation of cellulose-degrading bacteria and determination of their cellulolytic potential. Int. J. Microbiol. 2012; 2012, 578925.
JL Balcázar, ID Blas, I Ruiz-Zarzuela, D Vendrell, O Girones and JL Muzquiz. Sequencing of variable regions of the 16S rRNA gene for identification of lactic acid bacteria isolated from the intestinal microbiota of healthy salmonids. Comp. Immunol. Microbiol. Infect. Dis. 2007; 30, 111-8.
JS Johnson, DJ Spakowicz, BY Hong, LM Petersen, P Demkowicz, L Chen, SR Leopold, BM Hanson, HO Agresta, M Gerstein, E Sodergren and GM Weinstock. Evaluation of 16S rRNA gene sequencing for species and strain-level microbiome analysis. Nat. Commun. 2019; 10, 5029.
M Drancourt, C Bollet, A Carlioz, R Martelin, JP Gayral and D Raoult. 16S ribosomal DNA sequence analysis of a large collection of environmental and clinical unidentifiable bacterial isolates. J. Clin. Microbiol. 2000; 38, 3623-30.
M Wenzel, I Schönig, M Berchtold, P Kämpfer and H König. Aerobic and facultatively anaerobic cellulolytic bacteria from the gut of the termite Zootermopsis angusticollis. J. Appl. Microbiol. 2002; 92, 32-40.
GM Mathew, YM Ju, CY Lai, DC Mathew and CC Huang. Microbial community analysis in the termite gut and fungus comb of Odontotermes formosanus: The implication of Bacillus as mutualists. FEMS Microbiol. Ecol. 2012; 79, 504-17.
HRK Ali, NF Hemeda and YF Abdelaliem. Symbiotic cellulolytic bacteria from the gut of the subterranean termite Psammotermes hypostoma Desneux and their role in cellulose digestion. AMB Express 2019; 9, 111.
H König. Bacillus species in the intestine of termites and other soil invertebrates. J. Appl. Microbiol. 2006; 101, 620-7.
N Zhang, D Yang, JRA Kendall, R Borriss, IS Druzhinina, CP Kubicek, Q Shen and R Zhang. Comparative genomic analysis of Bacillus amyloliquefaciens and Bacillus subtilis reveals evolutional traits for adaptation to plant-associated habitats. Front. Microbiol. 2016; 7, 2039.
A Amore, O Pepe, V Ventorino, A Aliberti and V Faraco. Cellulolytic Bacillus strains from natural habitats-a review. Chim. Oggi 2013; 31, 49-52.
RK Rathnan, D John and T Balasaravanan. Isolation, screening, identification and optimized production of extracellular cellulase from Bacillus subtilis using cellulosic waste as carbon source. J. Microbiol. Biotechnol. Food Sci. 2013; 2, 2383-6.
Downloads
Published
How to Cite
Issue
Section
License
Copyright (c) 2021 Walailak University

This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License.