Real-Time Prediction of the COVID-19 Epidemic in Thailand using Simple Model-Free Method and Time Series Regression Model
DOI:
https://doi.org/10.48048/wjst.2021.10028Keywords:
COVID-19, Predictive model, Model-free method, Time series regression, Genetic algorithmAbstract
The novel coronavirus 2019 (COVID-19) pandemic was declared a global health crisis. The real-time accurate and predictive model of the number of infected cases could help inform the government of providing medical assistance and public health decision-making. This work is to model the ongoing COVID-19 spread in Thailand during the 1st and 2nd phases of the pandemic using the simple but powerful method based on the model-free and time series regression models. By employing the curve fitting, the model-free method using the logistic function, hyperbolic tangent function, and Gaussian function was applied to predict the number of newly infected patients and accumulate the total number of cases, including peak and viral cessation (ending) date. Alternatively, with a significant time-lag of historical data input, the regression model predicts those parameters from 1-day-ahead to 1-month-ahead. To obtain optimal prediction models, the parameters of the model-free method are fine-tuned through the genetic algorithm, whereas the generalized least squares update the parameters of the regression model. Assuming the future trend continues to follow the past pattern, the expected total number of patients is approximately 2,689 - 3,000 cases. The estimated viral cessation dates are May 2, 2020 (using Gaussian function), May 4, 2020 (using a hyperbolic function), and June 5, 2020 (using a logistic function), whereas the peak time occurred on April 5, 2020. Moreover, the model-free method performs well for long-term prediction, whereas the regression model is suitable for short-term prediction. Furthermore, the performances of the regression models yield a highly accurate forecast with lower RMSE and higher R2 up to 1-week-ahead.
HIGHLIGHTS
- COVID-19 model for Thailand during the first and second phases of the epidemic
- The model-free method using the logistic function, hyperbolic tangent function, and Gaussian function applied to predict the basic measures of the outbreak
- Regression model predicts those measures from one-day-ahead to one-month-ahead
- The parameters of the model-free method are fine-tuned through the genetic algorithm
GRAPHICAL ABSTRACT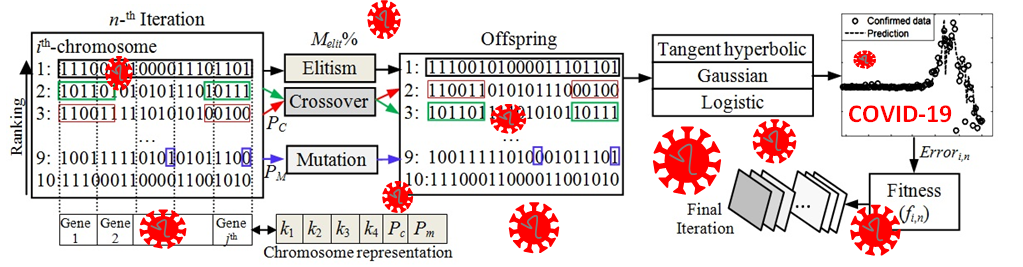
Downloads
Metrics
References
X Tan, E Feng and G Xu. SARS epidemic modeling and the study on its parameter control system. J. Eng. Math. 2003; 20, 39-40.
Y Bai and Z Jin. Prediction of SARS epidemic by BP neural networks with online prediction strategy. Chaos Solitons Fractals 2005; 26, 559-69.
A Earnest, MI Chen, D Ng and LY Sin. Using autoregressive integrated moving average (ARIMA) models to predict and monitor the number of beds occupied during a SARS outbreak in a teritiary hospital in Singapore. BMC Health Serv. Res. 2005; 5, 36.
OB Da’ar and AE Ahmed. Underlying trend, seasonality, prediction, forecasting and the contribution of risk factors: An analysis of globally reported cases of Middle East Respiratory Syndrome Coronavirus. Epidemiol. Infect. 2018; 146, 1343-49.
K Nah, S Otsuki, G Chowell and H Nishiura. Predicting the international spread of Middle East respiratory syndrome (MERS). BMC Infect. Dis. 2016; 16, 356.
WO Kermack and AG McKendrick. Contributions to the mathematical theory of epidemics - I. Bull Math. Biol. 1991; 53, 33-55.
W Yang, A Karspeck and J Shaman. Comparison of filtering methods for the modeling and retrospective forecasting of influenza epidemics. PLoS Comput. Biol. 2004; 10, e1003583.
TK Yamana, S Kandula and J Shaman. Individual versus superensemble forecasts of seasonal influenza outbreak in the United States. PLoS Comput. Biol. 2017; 13, e1005801.
EL Ray and NG Reich. Prediction of infectious disease epidemics via weighted density ensembles. PLoS Comput. Biol. 2018; 14, e1005910.
TM Chen, J Rui, QP Wang, ZY Zhao, JA Cui and L Yin. A mathematical model for simulating the phase-based transmissibility of a novel coronavirus. Infect. Dis. Poverty 2020; 9, 24.
Q Li, X Guan, P Wu, X Wang, L Zhou, Y Tong, R Ren, KSM Leung, EHY Lau, JY Wong, X Xing, N Xiang, Y Wu, C Li, Q Chen, D Li, T Liu, J Zhao, M Liu, W Tu, C Chen, L Jin, R Yang, Q Wang, S Zhou, R Wang, H Liu, Y Luo, Y Liu, G Shao, H Li, Z Tao, Y Yang, Z Deng, B Liu, Z Ma, Y Zhang, G Shi, TTY Lam, JT Wu, GF Gao, BJ Cowling, B Yang, GM Leung and Z Feng. Early transmission dynamics in Wuhan, China, of novel coronavirus-infected pneumonia. N. Eng. J. Med. 2020; 382, 1199-207.
CY Yang and J Wang. A mathematical model for the novel coronavirus epidemic in Wuhan, China. Math. Biosci. Eng. 2020; 17, 2708-24.
J Gu, Z Gao and W Li. Modeling of epidemic spreading with white Gaussian noise. Chinese Sci. Bull. 2011; 56, 3683-88.
G Zhou and G Yan. Severe acute respiratory syndrome epidemic in Asia. Emerg. Infect. Dis. 2003; 9, 1608-10.
YH Hsieh, JY Lee and HL Chang. SARS epidemiology modeling. Emerg. Infect. Dis. 2004; 10, 1165-7.
K Roosa, Y Lee, R Luo, A Kirpich, R Rothenberg, JM Hyman, P Yang and G Chowell. Short-term forecasts of the COVID-19 epidemic in Guangdong and Zhejiang, China: February 13-23, 2020. J. Clin. Med. 2020; 9, 596.
B Pirouz, SS Haghshenas, SS Haghshenas and P Piro. Investigating a serious challenge in the sustainable development process: Analysis of confirmed cases of COVID-19 (new type of coronavirus) through a binary classification using artificial intelligence and regression analysis. Sustainability 2020; 12, 2427.
Downloads
Published
How to Cite
Issue
Section
License
Copyright (c) 2021 Walailak University

This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License.